CORALL RNA-Seq V2 -
Your Key to Uncovering RNA Therapeutics Potential
Evaluate the activity of RNA therapeutics with CORALL RNA-Seq V2!

Time Saving
CORALL offers a fast and fully automatable whole transcriptome RNA-Seq workflow which is completed in only 4.5 hours.

Excels with Low Input
CORALL supports the widest input range (1 ng - 1000 ng) with excellent performance for low input RNA down to 1 ng prior to mRNA-selection or ribo-depletion.

Adjustable Size
CORALL RNA-Seq V2 allows fragmentation-free library size adjustment to fit your application. Choose between medium size for expression analysis or longer lengths for annotation-based applications.
![]()
Upscale Experiments with Ease
Automation-friendly and fast workflows allow processing of hundreds of samples in a single day and speed up turnover time to results!
Get the best sequencing results for demanding applications with CORALL RNA-Seq V2.
Add extra-confidence with built-in Unique Molecular Identifiers (UMIs) and process any sample type and quality with ease in only 6 steps: from sample to sequencing-ready library in less than one day.
Fast and flexible library prep in only 4.5 hours ⏱️
CORALL does not require RNA fragmentation and is thus perfectly suited for processing of degraded or FFPE RNA.

Figure 1 | Comparison of CORALL and conventional WTS RNA-Seq library preparation workflows
CORALL Workflow

Samples are treated using a set of affinity probes for specific depletion of rRNA sequences and magnetic beads for depletion. RiboCop efficiently removes rRNA while maintaining unbiased transcript expression.


CORALL library generation is initiated by random hybridization of Displacement Stop Primers (DSP) to the RNA template. The DSP also contains a partial Illumina adapter sequence. Reverse transcription extends each DSP to the next one, where transcription is effectively stopped.

Step 2: CORALL Library Generation: Linker Ligation.
Highly efficient ligation of Linker Oligos to the 3’ ends of first-strand cDNA fragments then introduces partial Illumina-adapters and Unique Molecular Identifiers (UMIs).


The UMI is positioned to be read out at the beginning of Read 1 during sequencing and can therefore be assessed even in Single-Read mode. All purification steps are based on magnetic beads, rendering the complete workflow highly suitable for automation.
Transcriptome-wide evaluation of siRNA activity with CORALL
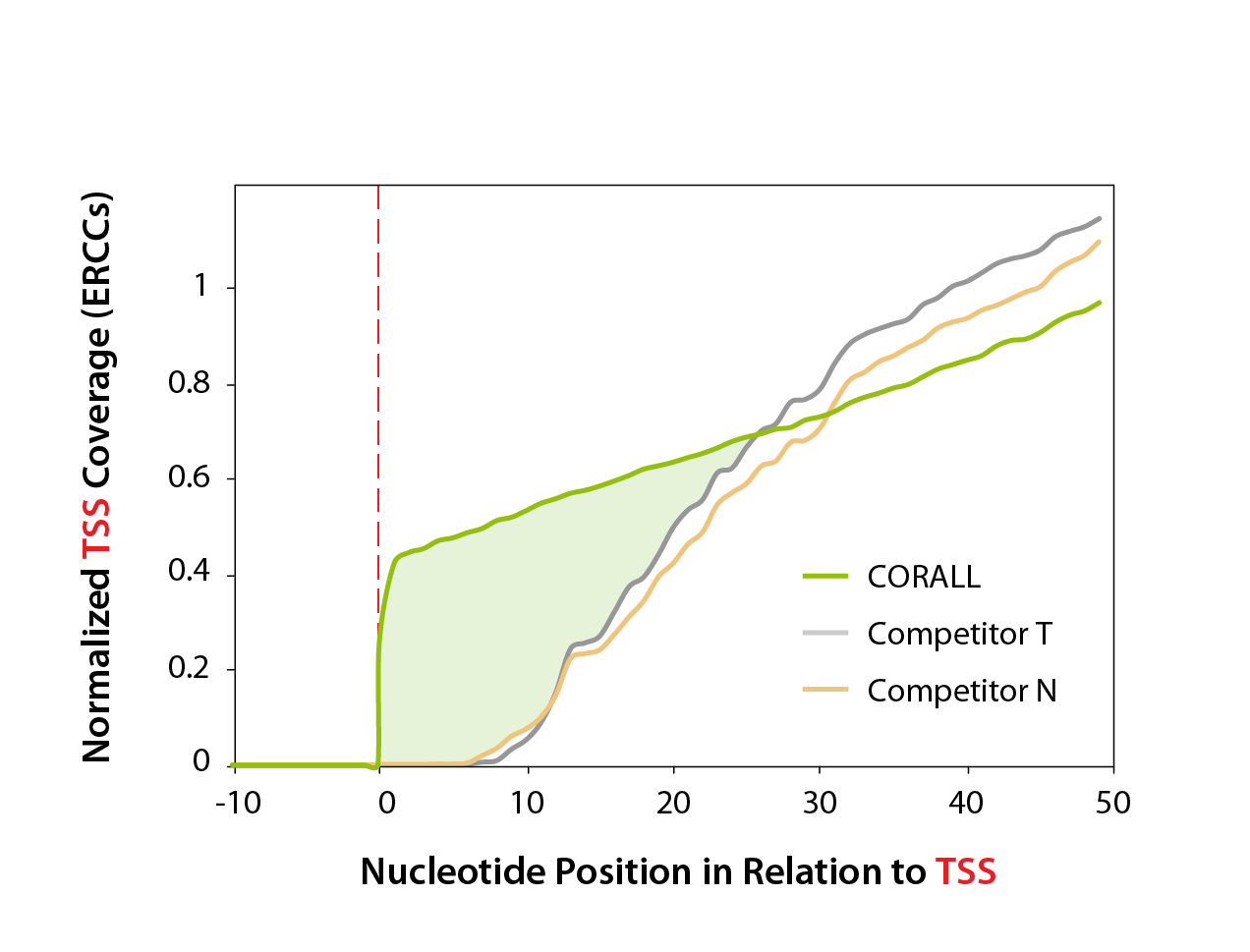
Figure 2 | Normalized ERCC coverage of transcription start sites. Normalized coverage of accumulated mapped reads for all detected ERCCs. The absolute nucleotide positions relative to the TSS (red dotted line) are shown.
Excellent start site representation
transcription start site representation. Read coverage was analyzed using the ERCC spike-in controls, which feature precise, known transcription start sites (TSS).
Validation of anti-fibrotic siRNAs therapeutics
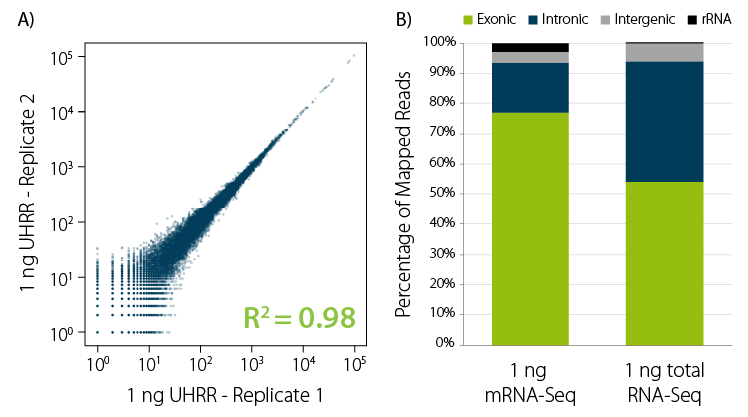
Figure 3 | Exceptional performance on low input RNA. A) Excellent correlation for replicates of 1 ng Universal Human Reference RNA (UHRR) processed with CORALL mRNA-Seq V2. B) Read distribution for 1 ng input RNA for CORALL mRNA-Seq V2 and CORALL Total RNA-Seq V2 with RiboCop rRNA depletion.
Excellent performance on low input RNA
Wide input range and exceptional sensitivity
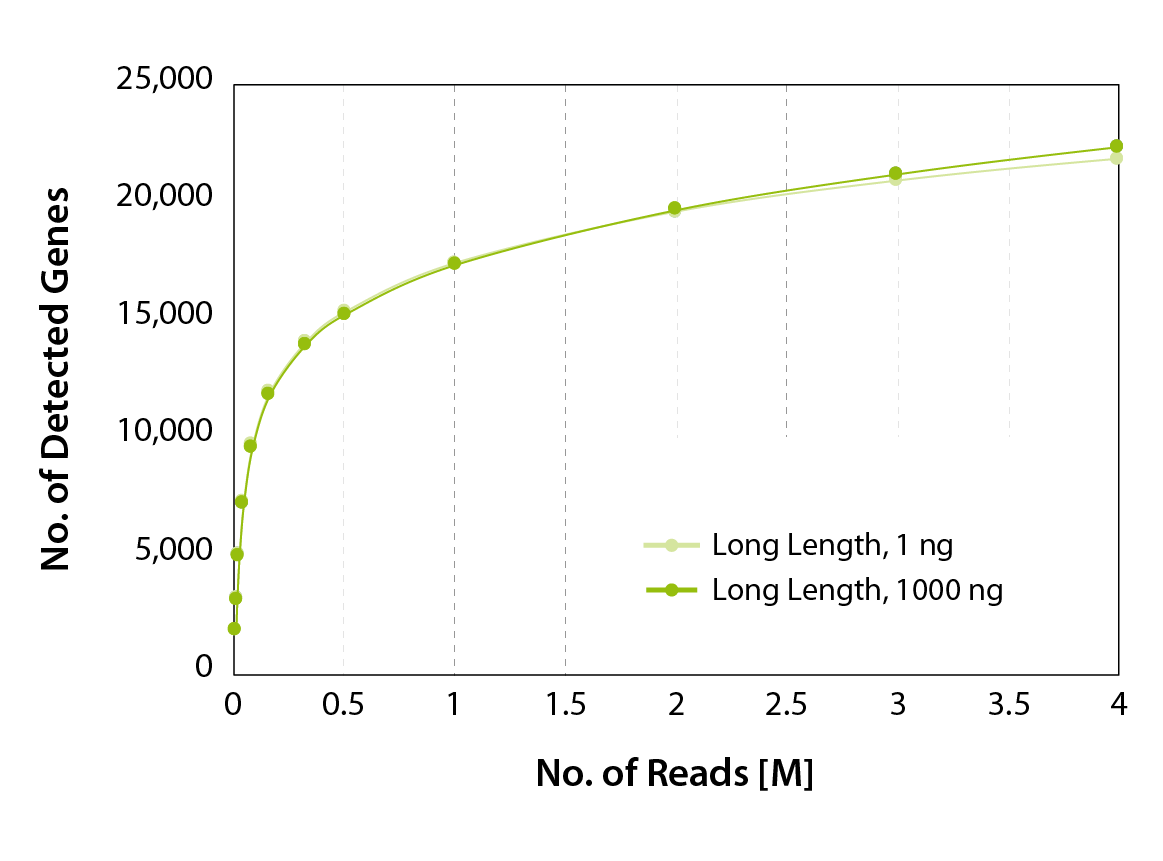
Figure 4 | Gene discovery rates. CORALL mRNA-Seq libraries with long inserts were preprared from 1000 ng and 1 ng UHRR . The number of detected genes was plotted against the total number of uniquely mapping exonic reads.

Upscale your experiments with ease!
CORALL RNA-Seq V2 Kits
CORALL RNA-Seq V2
-
Fragmentation-free RNA-Seq library prep with superior end-to-end coverage and true start site representation to cover all bases.
-
Built-in UMIs for highly accurate expression analysis
of coding and non-coding transcripts. -
CORALL RNA-Seq V2 supports the widest input range (1 ng - 1000 ng) with excellent sensitivity for low input RNA.
- Easily scalable automation-friendly workflow.

Ordering Information
-
Cat. № | 171 - 176
Product Name CORALL RNA-Seq V2 Library Prep Kit with UDIs (Set A1, A2, A3, A4, B1, or A1- A4)
Kit Sizes [preps] 24, 96, 384 -
Cat. № | 177 - 182
Product Name CORALL mRNA-Seq V2 Library Prep Kit with UDIs (Set A1, A2, A3, A4, B1, or A1- A4)
Kit Sizes [preps] 96, 384 -
Cat. № | 183 -184
Product Name RiboCop HMR V2 and CORALL Total RNA-Seq V2 Library Prep Kit with UDIs (A1 or B1)
Kit Sizes [preps] 24, 96 -
Cat. № | 185 - 186
Product Name RiboCop HMR+Globin and CORALL Total RNA-Seq V2 Library Prep Kit with UDIs (A1 or B1)
Kit Sizes [preps] 24, 96
References
2Ahn et al., 2022 Sci. Rep. (11): 19161, doi: 10.1038/s41598-021-98708-z
About Lexogen
Lexogen is a biotech company focusing on RNA and complete transcriptome studies using next generation sequencing technologies.
We have been at the forefront of RNA research since 2007, and our technologies and products are being used by thousands of scientists all over the world. “Lexo” in Greek means “word”, and Lexogen stands for the word of genes, which is the transcriptome. We strive to be the pioneers in this fascinating field of RNA science, and together with the fastest-growing technology of the past years, the next generation sequencing (NGS), we are sure that we can make a big contribution to the world.

